Media Matters 2: New Developments in Hematopoietic Cell Culture for Gene Editing and Cell Therapy Research

Update: Culturing HSPCs in Serum-Free Media for Cell Therapy Research
Life-saving treatments for a variety of diseases, particularly those that affect the blood and immune systems, depend on hematopoietic stem cells (HSCs).
In a previous post, we shone a spotlight on research breakthroughs and new methods for gene editing hematopoietic stem and progenitor cells (HSPCs), which consist of cells with both multilineage (stem cells) and restricted lineage (progenitor) potential. The focus then was on methods that employ CRISPR-based gene editing systems to produce genetically-modified, autologous HSPCs suitable for transplantation. In the two years since, several new protocols to genetically modify HSPCs have been published from groups at the forefront of this research.
Notably, recent research has seen a focus on standardizing protocols for gene editing HSPCs, using optimized culture conditions. There is increased recognition that, to successfully genetically modify HSPCs, culture conditions must both maintain the balance of stimulatory signals for proliferation and promote the properties that characterize ‘stemness’, such as self-renewal and multilineage differentiation potential. In this update, we highlight the impact of flexible and specific culture conditions on the recent developments in hematopoietic cell therapy research.
Traditionally, HSPCs have been cultured in liquid growth media supplemented with cytokines. To enhance HSPC expansion in vitro, culture media is now often augmented with the inclusion of novel small molecules. One such small molecule, UM171, stimulates in vitro expansion of human HSPCs through the upregulation of specific genes encoding for surface molecules, thereby enhancing the activity of the cell’s self-renewal machinery [1]. For example, endothelial protein C receptor (EPCR) expression in HSPCs is induced by UM171 treatment and has been shown to be a highly reliable surface marker for expanded human HSPCs [2].
Culture media for in vitro expansion and genome editing of HSPCs must perform reliably and consistently; ideally, they are also defined in their composition and optimized for the cell type of choice. StemSpanâ„¢ media contain no serum or cytokines, allowing researchers the flexibility to establish specific conditions for their experiment with their choice of expansion supplements, cytokines, and other additives–including small molecules.
The utility of a flexible media formulation is demonstrated by findings from studies employing specific media and supplement combinations. For example, when combined with cytokine supplements and UM171, StemSpanâ„¢ SFEM II medium promotes more than 10-fold expansion of primitive CD34+CD45RA–CD90+ stem/progenitor cells in serum-free conditions (Figure 1).
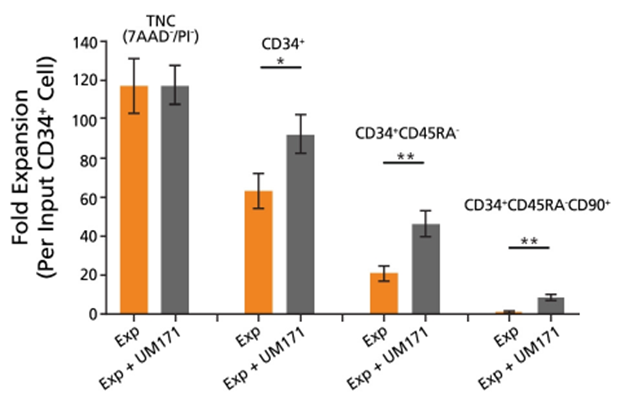
In 2018, Bak et al described a protocol for reproducible gene editing of human CD34+ cells [3]. Prior to gene editing, the CD34+ cells were cultured in a cytokine-rich, StemSpanâ„¢-based medium with added supplements and UM171. These specific conditions help to preserve ‘stemness’ while also promoting both cycling and survival of the cells in vitro. Elements of this protocol were described by Dr. Daniel Dever in a recent webinar.
Similarly, a 2018 study by Charlesworth et al [4] showed that low-density culture conditions using StemSpanâ„¢ SFEM II medium supplemented with UM171 and another small molecule known to promote HSPC expansion, StemRegenin 1 (SR1), enabled expansion of gene-edited HSPCs including, notably, the rare gene-edited long-term repopulating (LT)-HSCs.
Optimization of Gene Editing Methods
In many ways, gene editing is still in its infancy and current techniques, which can be cumbersome and expensive, continue to be developed and refined. However, this is an area that holds a great deal of promise for the investigation of the basic biology of various diseases and the development of new therapies. Here, we review recent research directed toward optimizing and perfecting gene editing techniques for HSPCs.
In 2017, Chicaybam et al developed a protocol for efficient electroporation of primary human and murine CD34+ cells using ‘Chicabuffers’ produced in their own laboratory [5]. The colony-forming unit (CFU) assay was used to evaluate hematopoietic progenitor cell function following electroporation. The result of their CFU assays suggested that their unique electroporation conditions preserve the function of myeloid, erythroid and granulocytic progenitor cells. Furthermore, the authors standardized the quantification of their CFU assays across multiple experiments by using STEMvisionâ„¢ for automated imaging and counting of hematopoietic colonies.
More recently, in 2019 Shahbazi et al [6] investigated the utility of a CRISPR nanoformulation based on gold nanoparticles, which entirely eliminates the need for either electroporation or viruses. Gold nanoparticles were found to localize in the nucleus and effectively escaped HSPC lysosomes, possibly due to the coating of polyethylenimine creating a proton sponge effect. Though further testing is required, this study indicates that gold nanoparticles have no adverse effect on cell viability and effectively delivered CRISPR gene editing machinery to primary HSPCs at levels comparable to electroporation.
Importantly, the use of culture media containing either bovine or human serum reduced total editing and homology-directed repair (HDR) rates with gold nanoparticles. This may be due to the formation of albumin corona by nanoparticles in the presence of serum albumin, which can alter their stability and uptake by cells [6]. This finding supports the importance of using serum-free media to support HSPC culture and gene editing.
Reproducibility with StemSpanâ„¢
In addition to improving conditions for gene editing, the use of serum-free media eliminates undefined elements from the critical pre- or post-electroporation phases of the culture system. These and other approaches toward standardization serve to increase the reproducibility of new gene editing techniques and protocols in labs around the world.
StemSpanâ„¢-ACF, the first commercially available animal component-free (ACF) culture medium for HSPCs, was used by Haase et al to generate ACF and transgene-free human induced pluripotent stem (hiPS) cells from CB CD34+ cells, which were found to express typical pluripotency markers with no residual Sendai virus (SeV) genome or transgene expression [7].
Using the same medium, in 2017 Agudelo et al designed a robust, marker-free co-selection strategy using both nuclease-driven nonhomologous end joining (NHEJ) and HDR gene editing [8]. The group used CRISPR-Cas9 and Cpf1/Cas13a systems to co-selected for dominant alleles of the sodium/potassium pump (Na+/K+ ATPase) and mechanistically related editing events at a second locus of interest. Cells transfected with CRISPR components were cultured with ouabain, a plant-derived inhibitor of Na+/K+ ATPase, in StemSpanâ„¢-ACF medium. Cells engineered to be resistant to ouabain were found to have successfully completed both genomic manipulations.
Using StemSpanâ„¢SFEM, in 2018 Ting et al were able to identify both known and previously unknown regulators of CD34+ HSPC expansion with their novel “Guide Swap” platform [9]. The development of this “Guide Swap” platform enables genome-scale pooled CRISPR-Cas9 screening in human primary cells, an approach which is unavailable with standard CRISPR-Cas9 gene editing techniques due to the requirement for stable and clonal Cas9-expressing cells.
Tools for Cell Therapy Research for Hematological Diseases and Other Disorders
To date, gene editing has shown promise for the treatment of a variety of diseases, such as β-hemoglobinopathies, including sickle cell anemia caused by mutations in the ɑ-globin and β-globin genes [11, 15] and has even been applied to engineer HIV resistance [12].
For example, De Ravin et al showed that stimulating HSPCs in StemSpanâ„¢ SFEM prior to electroporation with either zinc finger nucleases or CRISPR-Cas9, followed by template DNA delivery with an AAV6 donor, was effective in facilitating precise gene editing of HSPCs [10,11]. These techniques may be used for treatment of X-linked chronic granulomatous disease (X-CGD) and other monogenic diseases in the future.
CRISPR-Cas9 gene editing may also be used to mimic natural genetic mutations and, thus, employed as an alternative method of gene therapy. For example, Mettananda et al used CRISPR-Cas9 gene editing to reduce excess É‘-globin expression in human HSPCs, mimicking a natural mutation that reduces É‘-globin expression to a level shown to be beneficial to patients with ð›ƒ-thalassemia [12].
Using a similar approach, Xu et al targeted the CCR5 gene in LT-HSCs [13]. A naturally occurring mutation in CCR5 has previously been shown to confer a loss of detectable viral load in patients with HIV, indicating that mimicking this may be a viable treatment for HIV-1 infection [14]. CCR5 gene-edited LT-HSCs were found to reconstitute rapidly and effectively in immunodeficient NOD/Prkdcscid/IL-2Rγnull (NPG) mice, with no loss of in vitro colony-forming potential, and successfully conferred resistance to HIV-1 with minimal off-target effects. These CCR5-ablated LT-HSCs engrafted successfully and were detectable in mice for more than one year following transplantation [13].
Martyn et al [15] used the CRISPR-Cas9 genome editing system to investigate the underlying operational mechanism of hereditary persistence of fetal hemoglobin (HPFH), a rare, benign condition. They concluded that genetic mutations at specific sites of the ð›¾-globin gene promoter disrupt the binding of two major fetal globin repressors, BCL11A and ZBTB7A, possibly laying the foundation for genetic treatment of this syndrome via the introduction of naturally occurring variants.
Currently, many hematological disorders, including ð›ƒ-thalassemia, are managed through supportive treatments such as blood transfusions. However, the lack of a sustainable blood supply creates a need for an alternate, reliable source of red blood cells (and other cells). Trakarnsanga et al (2017) [16] presented a robust and reproducible gene editing method to generate an immortalized adult erythroid cell line by transducing primary bone marrow CD34+ HSPCs with a lentiviral vector before expanding in StemSpanâ„¢ SFEM II medium.
In summary, the past several years has seen several major advances in gene editing of HSPCs. These developments were facilitated in part by the use of standardized culture media and novel small molecules that help increase the proliferation of primitive stem cells while maintaining ‘stemness’. StemSpanâ„¢ media has been instrumental in many of gene editing advancements, by providing researchers with a flexible culture system that can be modified to meet researchers’ specific needs.
For more information and a step-by-step protocol on using StemSpanâ„¢ media, UM171, and other products for HSPC cell expansion, read our tech tip on the serum-free expansion of CD34+ cells with SFEM II and UM171. You can find more information about our tools for gene editing and watch our recent webinar with Daniel Dever from the Porteus lab, on our website.
References
- Fares I, Chagraoui J, Gareau Y et al. Cord blood expansion. Pyrimidoindole derivatives are agonists of human hematopoietic stem cell self-renewal. Science, 345(6203):1509-12 (2014)
- Fares I, Chagraoui J, Lehnertz B et al. EPCR expression marks UM171-expanded CD34+ cord blood stem cells. Blood, 129(25):3344-3351 (2017)
- Bak RO, Dever DP, Porteus MH. CRISPR/Cas9 genome editing in human hematopoietic stem cells. Nat. Protoc. 13(2): 358—376 (2018)
- Charlesworth CT, Camarena J, Cromer MK et al. Priming Human Repopulating Hematopoietic Stem and Progenitor Cells for Cas9/sgRNA Gene Targeting. Mol. Ther. Nucleic Acids, 12: 89—104 (2018)
- Chicaybam L, Barcelos C, Peixoto B et al. An Efficient Electroporation Protocol for the Genetic Modification of Mammalian Cells. Front. Bioeng. Biotechnol. 4: 99 (2017)
- Shahbazi R, Sghia-Hughes G, Reid JL et al. Targeted homology-directed repair in blood stem and progenitor cells with CRISPR nanoformulations. Nat. Mater. [Epub ahead of print] (2019)
- Haase A, Göhring G, Martin U. Generation of non-transgenic iPS cells from human cord blood CD34+ cells under animal component-free conditions. Stem Cell Res. 21: 71—73 (2017)
- Agudelo D, Duringer A, Bozoyan L et al. Marker-free coselection for CRISPR-driven genome editing in human cells. Nat. Methods, 14(6): 615—620 (2017)
- Ting PY, Parker AE, Lee JS et al. Guide Swap enables genome-scale pooled CRISPR-Cas9 screening in human primary cells. Nat. Methods, 15(11): 941—946 (2018)
- De Ravin SS, Reik A, Liu PQ et al. Targeted gene addition in human CD34(+) hematopoietic cells for correction of X-linked chronic granulomatous disease. Nat. Biotechnol. 34(4): 424—9 (2016)
- De Ravin SS, Li L, Wu X et al. CRISPR-Cas9 gene repair of hematopoietic stem cells from patients with X-linked chronic granulomatous disease. Sci. Transl. Med. 9(372) (2017)
- Mettananda S, Fisher CA, Hay D et al. Editing an α-globin enhancer in primary human hematopoietic stem cells as a treatment for β-thalassemia. Nat. Commun. 8(1): 424 (2017)
- Xu L, Yang H, Gao Y et al. CRISPR/Cas9-Mediated CCR5 Ablation in Human Hematopoietic Stem/Progenitor Cells Confers HIV-1 Resistance In Vivo. Mol. Ther. 25(8): 1782—1789 (2017)
- Allers K, Hütter G, Hofmann J et al. Evidence for the cure of HIV infection by CCR5Δ32/Δ32 stem cell transplantation. Blood, 117(10): 2791—9 (2011)
- Martyn GE, Wienert B, Yang L et al. Natural regulatory mutations elevate the fetal globin gene via disruption of BCL11A or ZBTB7A binding. Nat. Genet. 50(4): 498—503 (2018)
- Trakarnsanga K, Griffiths RE, Wilson MC et al. An immortalized adult human erythroid line facilitates sustainable and scalable generation of functional red cells. Nat. Commun. 8: 14750 (2017)